Abstract
Single Cell Sequencing is an emerging technique in the field of Next Generation Sequencing (NGS). Starting with the RNA isolated from a single cell, there is a big need for high quality library preparation and size selection methods. Sample purification is an important part of the library preparation. This experiment describes the use of CleanNGS1) magnetic beads offered by CleanNA1) to do these purifications in the library preparation for Single Cell Sequencing. The CleanNGS1) magnetic beads can simultaneously be used for the size selection steps in the library preparation. The conclusions from this experiment show that the prepared libraries have a high purity and high quality and meet all the demands for size selection.
Introduction
With emerging NGS technologies, for example like Single Cell Sequencing, there is an increased need for NGS library purification methods providing accurate results starting from low input amounts of DNA and/or RNA. This application note will demonstrate the abilities of CleanNGS to provide, high recovery and accurate size selection abilities. A Single Cell library series have been prepared to demonstrate the CleanNGS purification, recovery and size selection abilities.
Equipment:
- Agilent, BioAnalyzer™ 2100
- DeNovix®, DS-11 FX
- Permagen™ 96-well Magnetic Separation Plate (P/N S480) 1),2)
Labware:
- 15 mL Greiner tubes
- PCR plates
Chemicals:
- CleanNGS (P/N CNGS-0050)1),2)
- DeNovix Broad Range Assay
- Ethanol absolute
- Nuclease free water
- Single Cell Library preparation enzymes and buffers
Single Cell Sequencing library preparation
A series of 4 Single Cell libraries were prepared in duplicate, using CleanNGS in one and competitor A in the other duplicate series for the cleanups and size selection.
The procedure for library preparation:
- FACS sorting, to obtain 1 single cell per well
- Single Cell lysis and annealing of the RT-primer
- Reverse transcription
- Pre-amplification
- Clean-up using CleanNGS
- Tagmentation
- Tagmentation enzyme inactivation
- Amplification PCR
- Clean-up using CleanNGS
- Library check via Agilent BioAnalyzer and DeNovix DS-11 FX.
Results
Yield and size of the samples after library preparation was compared to competitor A. Library fragment size was compared using Agilent Technologies BioAnalyzer, while final library concentration and yield were determined via fluorescence measurement using a DeNovix DS-11 FX and the Denovix Broad Range Assay. Average library size and the concentration of each library can be found in table 1.
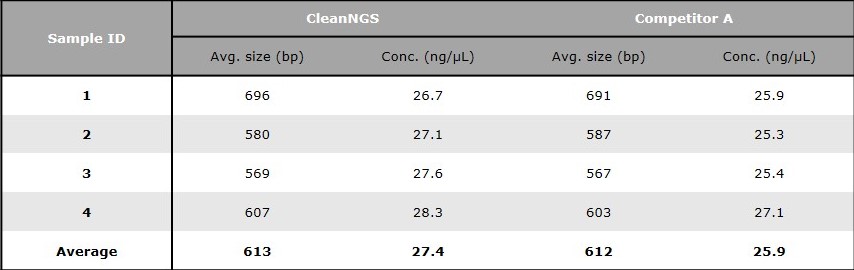
Table 1. Comparison of average library size and concentration between CleanNGS and competitor A.
The corresponding BioAnalyzer data can be found in figures 1 till 5, showing the electropherograms from samples processed with CleanNGS and competitor A.
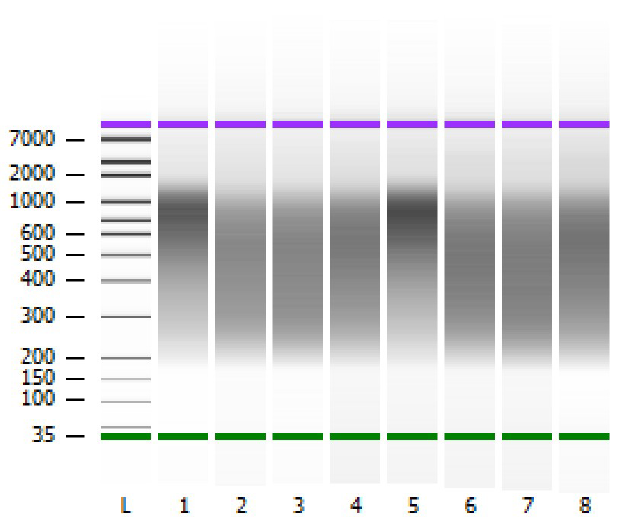
Figure 1. BioAnalyzer overview of the libraries. L=ladder, samples 1-4=competitor A, samples 5–8=CleanNGS.
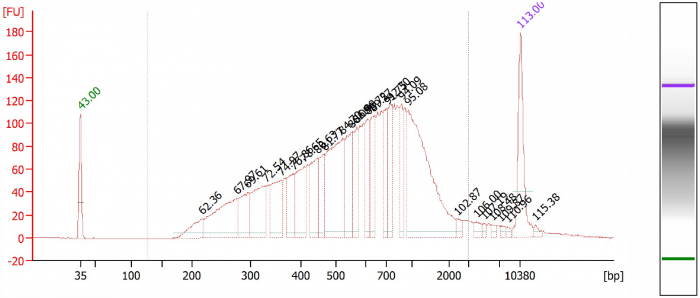
Figure 2. Library 1, competitor A.
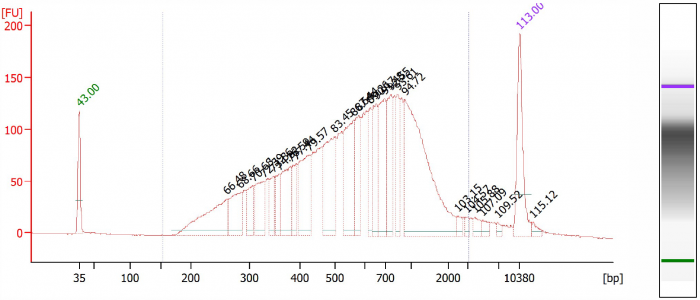
Figure 3. Library 1, CleanNGS
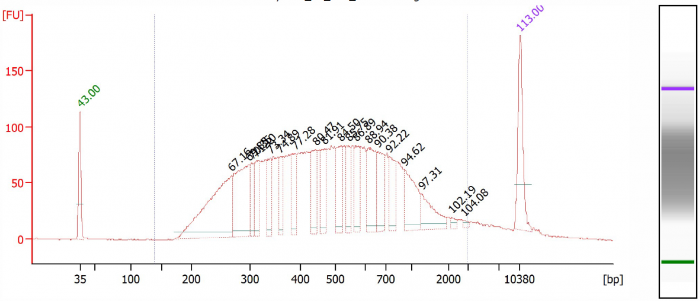
Figure 4. Library 2, competitor A.
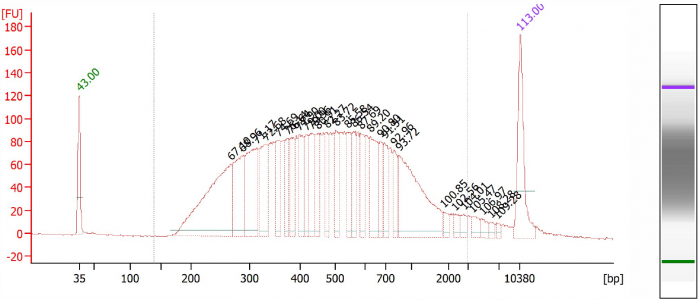
Figure 5. Library 2, CleanNGS.
Conclusion & Discussion
Both the preparation of the Single Cell Sequencing library as well as the qPCR show CleanNGS provides a solution for any NGS application. CleanNGS provides a high recovery and accurate size selection within the library prep processes. The size selection compared to competitor A, shows an average difference of 1 bp, which is considered to be the same. Especially taking into account the Agilent Bioanalyzer has a sizing resolution of ≥ 5 bp. Being well within this range with the size selection comparison shows both kits perform identical.
The recovery compared to competitor A as shown, has improved by an average of 5% on the samples used within this experiment. By improving the buffer composition of CleanNGS it binds DNA/RNA more effective compared to competitor A. The higher recovery makes CleanNGS a more powerful asset to be used in applications with even the lowest DNA/RNA inputs at the start of the experiment, by providing the best recovery of purified DNA/RNA.
Due to the protocol design, CleanNGS can be used both manually as well as automated. CleanNGS can be adopted in any NGS laboratory independent off sample throughput. Since the CleanNGS protocol is equal to competitor A, CleanNGS can be used as a direct replacement in already established laboratory processes.
To enable a broader usage of CleanNGS within NGS, but also in RNA applications such as MicroArrays, CleanNGS will be produced RNase free from the start of 2017. This will enhance performance of in vitro RNA applications with the goal of providing the best possible recovery within any in vitro RNA application.
References
- http://www.cleanna.com
- http://www.gcbiotech.com
- http://www.bioline.com
Ordering details CleanNGS
| Catalog No# | Description | Number of Reactions* |
| CNGS-0005 | CleanNGS (5 mL) | 277 |
| CNGS-0050 | CleanNGS (50 mL) | 2,777 |
| CNGS-0500 | CleanNGS (500 mL) | 27,777 |
Trademarks
Bio-Rad is a registered trademark of Bio-Rad Laboratories, Inc.
Bioline is a registered trademark of Meridian Life Science, Inc.
CFX96 Touch is a registered trademark of Bio-Rad Laboratories, Inc.
CleanNA is a registered trademark of GC biotech BV
CleanNGS is a registered trademark of GC biotech BV
Covaris is a registered trademark of Covaris, Inc.
Permagen is a registered trademark of Permagen Labware, LTD

